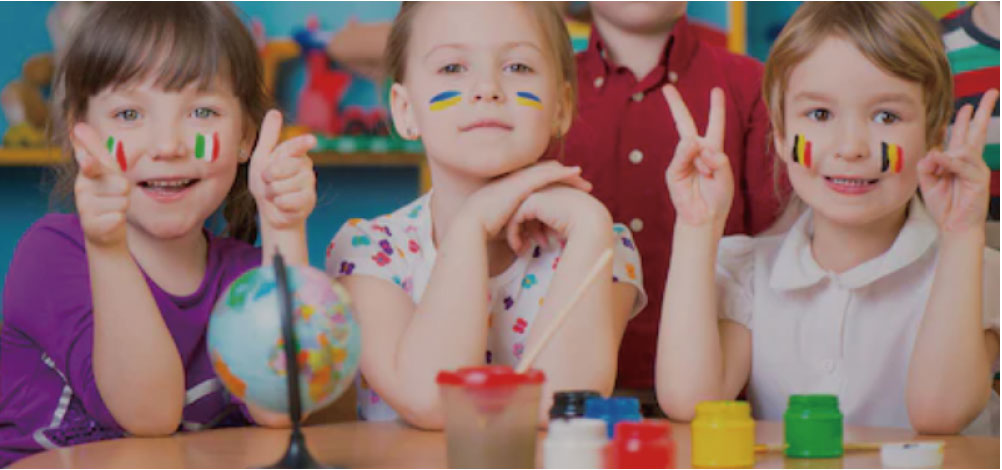
International “Holistic Education” for Global Citizens
― 国際的な真の「全人教育」 ―
国際バカロレアプログラムのPYP認定校として高度な知的水準と語学力、そして異文化に対する理解と寛容性を育むための「全人教育」を行っています。
0歳〜12歳までの幼小一貫教育を通じて、思考力・表現力・倫理能力・研究能力を育む教科横断的カリキュラムにより、物事を深く探求し、
多角的な考え方ができる
人物の育成を目指しています。
About “International Baccalaureate”
高い思考力・問題解決力を備え、
世界へはばたく人材を育成する

国際バカロレアが目指す学習者像の10項目
- Inquirers– 探究する人
- Knowledgeable– 知識のある人
- Thinkers– 考える人
- Communicators– コミュニケーションができる人
- Principled– 信念のある人
- Open-minded– 心を開く人
- Caring– 思いやりのある人
- Risk-takers– 挑戦する人
- Balanced– バランスのとれた人
- Reflective– 振り返りができる人
国際バカロレア(IB)とは、 「世界の複雑さを理解し、それに対処できる生徒」を育成するための
チャレンジに満ちた総合的な国際教育プログラムです。
教科横断型の多様なカリキュラムを通じて、高い語学力はもとより、責任ある行動をとるための「品性」と「知見」を習得し、
知的成長や学習面での成功にとどまらない、人間としての幅広い能力と責任感を育むことを目指す、国際的な全人教育を行っています。
Courses

生後57日〜5歳まで|週2/3/4/5回
『Prep.Prechool』はお子さまを生後57日からお預かりしております。また、『Preschool』では、基本となるWriting/Reading/Speaking/Listeningの学習とともに、各クラスごとのゴールに向かっての授業を行います。
Prep.Pre
&
Preschool

6歳~12歳まで|月曜〜金曜
「English」「Reading」「Writing」を中心とし、公立小学校と同等の算数、理科、社会、日本語の教育を実施。国際バカロレア理論に基づき、ディスカッションや社会見学、実習など、全人教育に寄与する総合科目も実施します。
Elementary

産休明け〜2歳まで|月曜〜土曜
一般社団法人アップビートインターナショナルスクールが運営する名古屋市認可小規模保育施設です。子どもの「何故?」を大切に、愛情をもって応え、導いていきます。
名古屋市認可
小規模保育園

幼児〜大人まで|クラス毎の開催日
生後6か月~1歳半までのBaby-Momクラス、プライベートレッスン、セミプライベートレッスンなど多種多様なコースをご用意しております。
Enrichment Courses

幼児~小学生まで|放課後・土曜日
アメリカ現地の教科書を使用し学年に合わせた英語力を伸ばすインター卒・帰国子女向けのAdvanced Course、また英語を基礎から学ぶBasic Courseを開講しています。
After
&
Saturday

小学1年〜6年まで|平日・土曜日・長期休業期間
留守家庭のお子様が安全に教育的で有意義な時間を過ごせるよう、UPBEATオリジナルの「多言語サポート」×「教育」×「食育」学童プログラムを提供いたします。
学童
Official Blog
 | 2024.04.12 3月28日~29日の1泊2日で、スプリングオーバーナイトキャンプを開催しました。 オーバーナイトキャンプって何? オーバーナイトキャンプはUP ... |
|---|---|
 | 2024.03.12 International Women’s Day (国際女性の日) 3月8日は、女性の権利遵守や社会的地位向上とジェンダー平等の実現を目指して制定されたInternational Women’s Day(国際女性の ... |
Contact us
入園・入学に関するご相談はお気軽にご相談ください。




















